Knowledge graph embedding for predicting and analyzing microbial interactions
Abstract
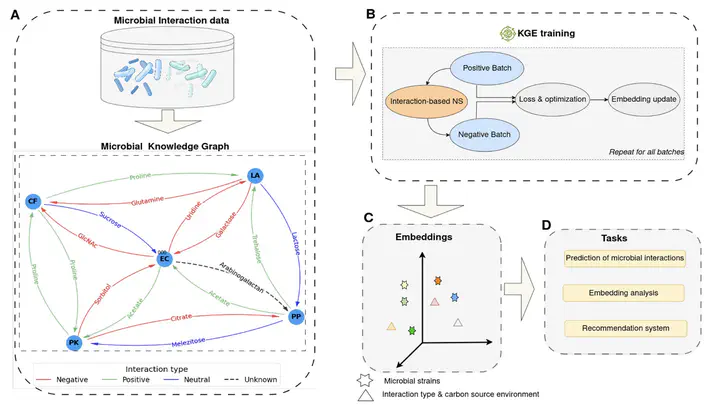
Interactions between microorganisms play a major role in shaping the structure and function of microbial communities, yet their prediction remains a challenge in microbial ecology. While currently available machine learning methods have shown promising performance, they often rely on extensive input features that are obtained from labor-intensive experiments. Here, we propose a new framework to predict pairwise interactions that minimizes the need for in vitro experimentation. Our approach is based on knowledge graph embedding, which learns the representation of microorganisms and their interactions in an embedding space. Using a dataset of interactions between 20 soil bacterial strains cocultured in 40 different carbon source environments, we demonstrate the effectiveness of our framework in accurately predicting pairwise interactions. Notably, we show that our model can predict interactions involving strains with missing culture data. We additionally show that the obtained embeddings can reveal similarities between carbon source environments, enabling the prediction of interactions in one environment based on the outcomes in a similar environment between the same pair of microorganisms. Furthermore, our approach allows the design of a recommendation system that can be used to guide microbial community engineering. These findings demonstrate that knowledge graph embedding is a promising modeling strategy in microbial ecology.